BioDeck Pipeline
Follow our streamlined computational workflow from data preparation to precise dynamic simulations.
Available Tools16
RDKit
Open-source cheminformatics software for molecule manipulation, descriptor calculation, and fingerprint generation.
Ligand Prep
Prepare small molecule ligands for docking simulations by correcting structures and generating 3D conformers.
Protein Prep
Prepare PDB receptor structures for docking — smart chain selection, solvent cleanup, metal ion preservation, and pH 7.4 protonation.

Sequence Crawler
Fetch DNA and Protein sequences from NCBI and UniProt using universal sequence crawling modules.
Auto GraphPad
Automated scientific data visualization. Upload CSV or Excel files to instantly generate publication-ready charts — no backend required.
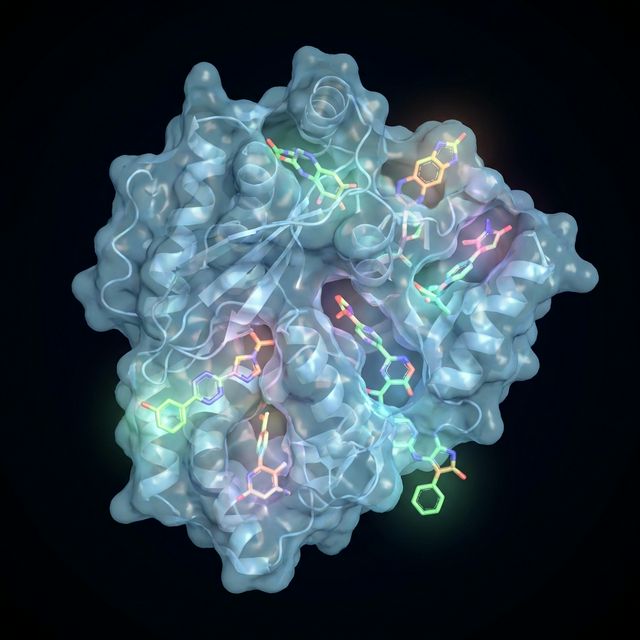
GNINA Screening
Deep learning-based molecular docking for virtual screening of large compound libraries against protein targets.
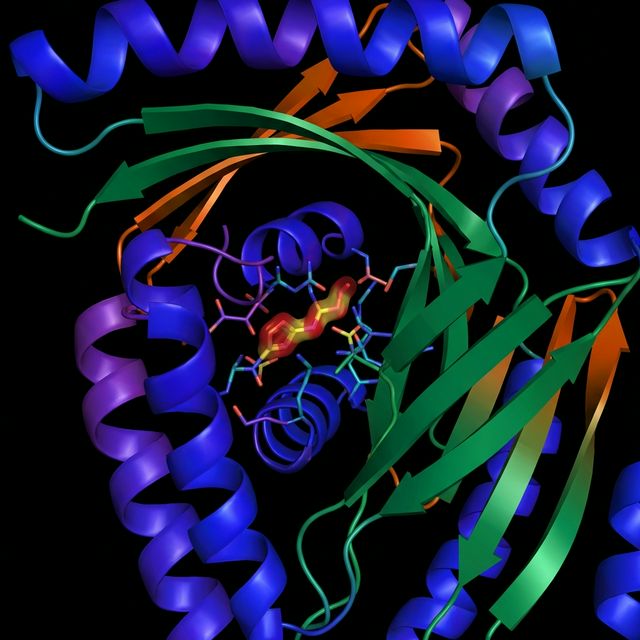
DiffDock-L
Latest iteration of diffusion-based docking for blind docking with high accuracy and improved sampling efficiency.
DynamicBind v2
Deep generative model for flexible protein-ligand docking with conformational sampling and animations.
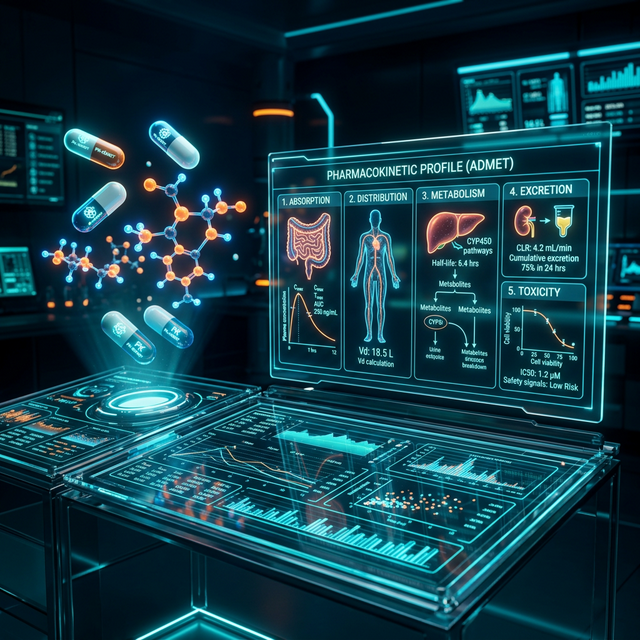

ADMET Screening
Predict ADMET properties swiftly and accurately using BioDeck.
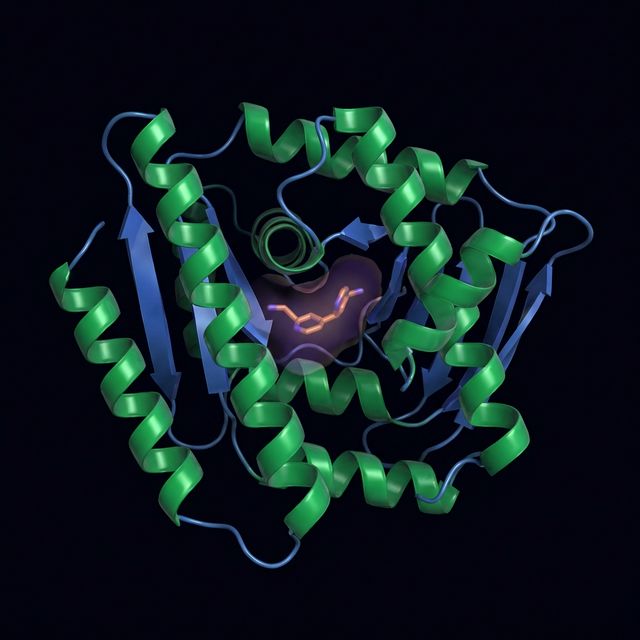
GNINA Optimization
Refine poses and scores of docked ligands using CNN scoring functions for precise binding affinity prediction.
Molecular Dynamics
Simulate the physical movements of atoms and molecules over time to understand complex biological systems.
MM-GBSA
Calculate the free energy of binding for protein-ligand complexes using molecular mechanics and generalized Born surface area continuum solvation.

RFdiffusion
Design novel protein structures, binders, and scaffolds using the deep learning model from RosettaCommons.

ProteinMPNN
Generate high-quality sequence designs for given protein backbones with thermodynamic stability.
BoltzGen
No-code YAML Builder for BoltzGen — design peptide, protein, and nanobody binders targeting specific residues with a generative AI.

ESM Phylo
Protein sequence phylogenetic tree analysis powered by ESM-C (EvolutionaryScale). Upload a FASTA file to generate embeddings, distance matrices, and interactive evolutionary trees.